Shikimate pathway
The shikimate pathway (shikimic acid pathway) is a seven-step metabolic pathway used by bacteria, archaea, fungi, algae, some protozoans, and plants for the biosynthesis of folates and aromatic amino acids (tryptophan, phenylalanine, and tyrosine). This pathway is not found in mammals.
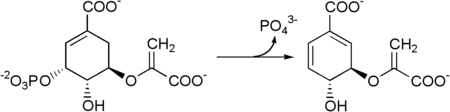
The seven enzymes involved in the shikimate pathway are DAHP synthase, 3-dehydroquinate synthase, 3-dehydroquinate dehydratase, shikimate dehydrogenase, shikimate kinase, EPSP synthase, and chorismate synthase. The pathway starts with two substrates, phosphoenol pyruvate and erythrose-4-phosphate, and ends with chorismate (chrorismic acid), a substrate for the three aromatic amino acids. The fifth enzyme involved is the shikimate kinase, an enzyme that catalyzes the ATP-dependent phosphorylation of shikimate to form shikimate 3-phosphate (shown in the figure below).[1] Shikimate 3-phosphate is then coupled with phosphoenol pyruvate to give 5-enolpyruvylshikimate-3-phosphate via the enzyme 5-enolpyruvylshikimate-3-phosphate (EPSP) synthase. Glyphosate, the herbicidal ingredient in Roundup, is a competitive inhibitor of EPSP synthase, acting as a transition state analog that binds more tightly to the EPSPS-S3P complex than PEP and inhibits the shikimate pathway.
Then 5-enolpyruvylshikimate-3-phosphate is transformed into chorismate by a chorismate synthase.
Prephenic acid is then synthesized by a Claisen rearrangement of chorismate by chorismate mutase.[2][3]
Prephenate is oxidatively decarboxylated with retention of the hydroxyl group to give p-hydroxyphenylpyruvate, which is transaminated using glutamate as the nitrogen source to give tyrosine and α-ketoglutarate.
References
- ^ Herrmann, K. M.; Weaver, L. M. (1999). "The Shikimate Pathway". Annual Review of Plant Physiology and Plant Molecular Biology. 50: 473–503. doi:10.1146/annurev.arplant.50.1.473. PMID 15012217.
- ^ Helmut Goerisch (1978). "On the mechanism of the chorismate mutase reaction". Biochemistry. 17 (18): 3700–3705. doi:10.1021/bi00611a004. PMID 100134.
- ^ Peter Kast; Yadu B. Tewari; Olaf Wiest; Donald Hilvert; Kendall N. Houk; Robert N. Goldberg (1997). "Thermodynamics of the Conversion of Chorismate to Prephenate: Experimental Results and Theoretical Predictions". J. Phys. Chem. B. 101 (50): 10976–10982. doi:10.1021/jp972501l.
Bibliography

- Edwin Haslam (1993). Shikimic Acid: Metabolism and Metabolites (1st ed.). ISBN 0471939994.
- Brown, Stewart A.; Neish, A. C. (1955). "Shikimic Acid as a Precursor in Lignin Biosynthesis". Nature. 175 (4459): 688–689. Bibcode:1955Natur.175..688B. doi:10.1038/175688a0. ISSN 0028-0836. PMID 14370198. S2CID 4273320.
- Weinstein, L. H.; Porter, C. A.; Laurencot, H. J. (1962). "Role of the Shikimic Acid Pathway in the Formation of Tryptophan in Higher Plants : Evidence for an Alternative Pathway in the Bean". Nature. 194 (4824): 205–206. Bibcode:1962Natur.194..205W. doi:10.1038/194205a0. ISSN 0028-0836. S2CID 4160308.
- Wilson, D J; Patton, S; Florova, G; Hale, V; Reynolds, K A (1998). "The shikimic acid pathway and polyketide biosynthesis". Journal of Industrial Microbiology and Biotechnology. 20 (5): 299–303. doi:10.1038/sj.jim.2900527. ISSN 1367-5435. S2CID 41117722.
- v
- t
- e
 Carbon fixation Photo- respiration Pentose phosphate pathway Citric acid cycle Glyoxylate cycle Urea cycle Fatty acid synthesis Fatty acid elongation Beta oxidation Peroxisomal beta oxidation
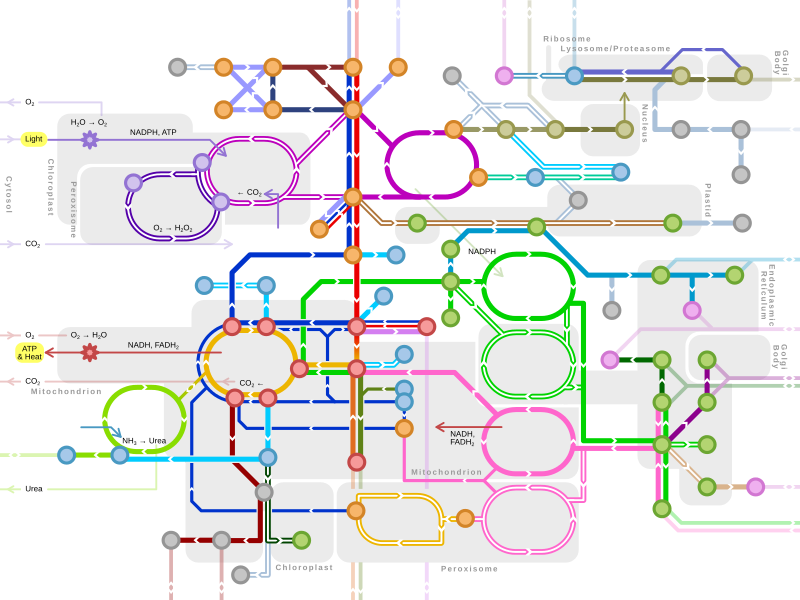
Glyco- genolysis Glyco- genesis Glyco- lysis Gluconeo- genesis Pyruvate decarb- oxylation Fermentation Keto- lysis Keto- genesis feeders to gluconeo- genesis Direct / C4 / CAM carbon intake Light reaction Oxidative phosphorylation Amino acid deamination Citrate shuttle Lipogenesis Lipolysis Steroidogenesis MVA pathway MEP pathway Shikimate pathway Transcription & replication Translation Proteolysis Glycosyl- ation
Sugar acids Simple sugars Propionyl -CoA Acetyl -CoA Acetyl -CoA Oxalo- acetate Succinyl -CoA α-Keto- glutarate Ketone bodies Respiratory chain Branched-chain amino acids Aspartate group Homoserine group & lysine Glutamate group & proline Ketogenic & glucogenic amino acids Aromatic amino acids & histidine Bile pigments Cobalamins (vitamin B12) Various vitamin Bs Cofactors Nucleic acids Acetyl -CoA Terpenoid backbones Glycero- phospholipids Fatty acids Glyco- sphingolipids Polyunsaturated fatty acids Endo- cannabinoids |
Single lines: pathways common to most lifeforms. Double lines: pathways not in humans (occurs in e.g. plants, fungi, prokaryotes).
 Orange nodes: carbohydrate metabolism.
Orange nodes: carbohydrate metabolism.  Violet nodes: photosynthesis.
Violet nodes: photosynthesis.  Red nodes: cellular respiration.
Red nodes: cellular respiration.  Pink nodes: cell signaling.
Pink nodes: cell signaling.  Blue nodes: amino acid metabolism.
Blue nodes: amino acid metabolism.  Grey nodes: vitamin and cofactor metabolism.
Grey nodes: vitamin and cofactor metabolism.  Brown nodes: nucleotide and protein metabolism.
Brown nodes: nucleotide and protein metabolism.  Green nodes: lipid metabolism.
Green nodes: lipid metabolism.