| BEX1 |
|---|
|
| Identifiers |
|---|
| Aliases | BEX1, BEX2, HBEX2, HGR74-h, brain expressed X-linked 1 |
|---|
| External IDs | OMIM: 300690; MGI: 1338017; HomoloGene: 135613; GeneCards: BEX1; OMA:BEX1 - orthologs |
|---|
| Gene location (Human) |
|---|
 | | Chr. | X chromosome (human)[1] |
|---|
| | Band | Xq22.1|Xq22 | Start | 103,062,651 bp[1] |
|---|
| End | 103,064,171 bp[1] |
|---|
|
| Gene location (Mouse) |
|---|
 | | Chr. | X chromosome (mouse)[2] |
|---|
| | Band | X F1|X 57.37 cM | Start | 134,967,313 bp[2] |
|---|
| End | 134,968,985 bp[2] |
|---|
|
| RNA expression pattern |
|---|
| Bgee | | Human | Mouse (ortholog) |
|---|
| Top expressed in | - endothelial cell
- Brodmann area 23
- middle temporal gyrus
- orbitofrontal cortex
- primary visual cortex
- dorsolateral prefrontal cortex
- superior frontal gyrus
- Brodmann area 9
- prefrontal cortex
- pituitary gland
|
| | Top expressed in | - arcuate nucleus
- ventromedial nucleus
- paraventricular nucleus of hypothalamus
- temporal lobe
- amygdala
- dorsomedial hypothalamic nucleus
- lateral septal nucleus
- anterior amygdaloid area
- lateral hypothalamus
- median eminence
|
| | More reference expression data |
|
|---|
| BioGPS | 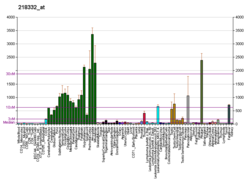 | | More reference expression data |
|
|---|
|
| Gene ontology |
|---|
| Molecular function | | | Cellular component | - cytoplasm
- nucleus
- transcription regulator complex
| | Biological process | - multicellular organism development
- cell differentiation
- positive regulation of DNA-binding transcription factor activity
- nervous system development
- positive regulation of transcription by RNA polymerase II
| | Sources:Amigo / QuickGO |
|
| Orthologs |
|---|
| Species | Human | Mouse |
|---|
| Entrez | | |
|---|
| Ensembl | | |
|---|
| UniProt | | |
|---|
| RefSeq (mRNA) | | |
|---|
| RefSeq (protein) | | |
|---|
| Location (UCSC) | Chr X: 103.06 – 103.06 Mb | Chr X: 134.97 – 134.97 Mb |
|---|
| PubMed search | [3] | [4] |
|---|
|
| Wikidata |
| View/Edit Human | View/Edit Mouse |
|


















